Structures
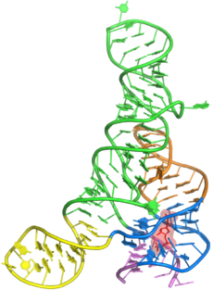
PRPP riboswitch
6DLT – Native
6DLR – Iridium hexammine soak
6DLQ – Mn2+ soak
6DLS – Tl+ soak
6DNR – apo structure
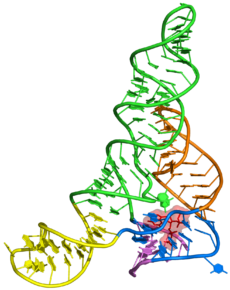
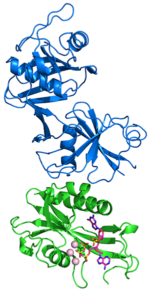
Crystal structure of RppH-DapF complex
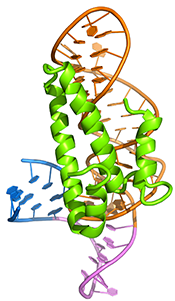
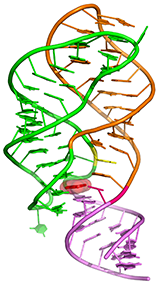
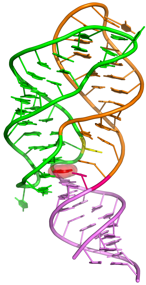
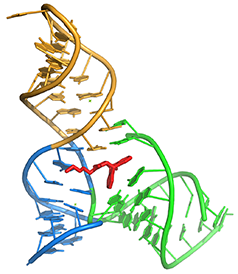
Diels-Alder ribozyme
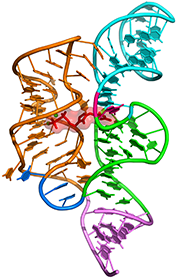
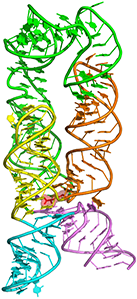
Lysine riboswitch
- 3DIS – Free form
- 3DIL – Bound to lysine
- 3DIO – Iridium hexammine soak
- 3DIM – Cs+ soak
- 3DIY – Mn2+ soak
- 3DJ2 – Tl+ soak
- 3DIX – K+ anomalous signal
- 3DIZ – Without Mg2+
- 3DIG – Bound to S-(2-aminoethyl)-L-cysteine
- 3DIQ – Bound to homoarginine
- 3DIR – Bound to N6-1-iminoethyl-L-Lysine
- 3DJ0 – Bound to L-4-oxalysine
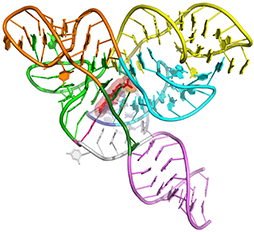
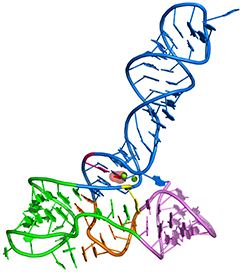
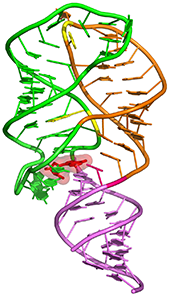
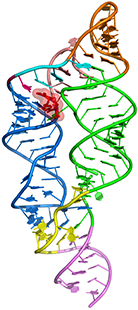
Tetrahydrofolate riboswitch
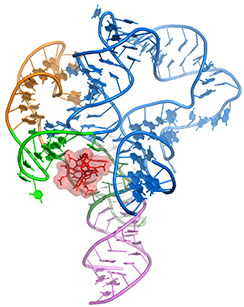
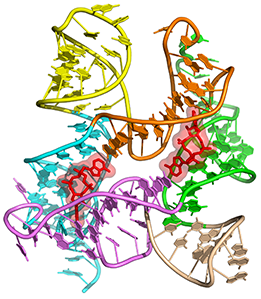
c-di-AMP riboswitch
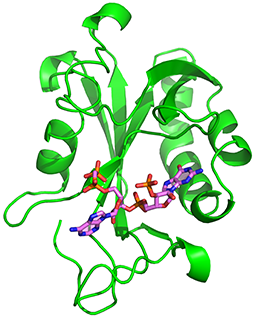
RppH bound to RNA and in free state
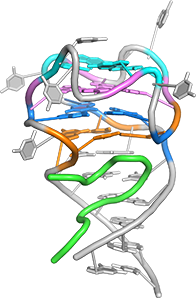
Complex between human FMRP RGG motif and G-quadruplex RNA

